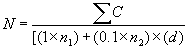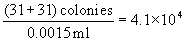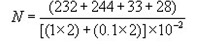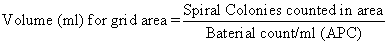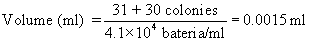BAM Chapter 3: Aerobic Plate Count
January 2001
Authors: Larry Maturin (ret.) and James T. Peeler (ret)
For additional information, contact Guodong Zhang.
The aerobic plate count (APC) is intended to indicate the level of microorganism in a product. Detailed procedures for determining the APC of foods have been developed by the Association of Official Analytical Chemists (AOAC) (3) and the American Public Health Association (APHA) (1). The conventional plate count method for examining frozen, chilled, precooked, or prepared foods, outlined below, conforms to AOAC Official Methods of Analysis, sec. 966.23, with one procedural change (966.23C). The suitable colony counting range (10) is 25-250. The automated spiral plate count method for the examination of foods and cosmetics (5), outlined below, conforms to AOAC Official Methods of Analysis, sec. 977.27. For procedural details of the standard plate count, see ref. 2.Guidelines for calculating and reporting plate counts have been changed to conform with the anticipated changes in the 16th edition of Standard Methods for the Examination of Dairy Products (2) and the International Dairy Federation (IDF) procedures (6).
Conventional Plate Count Method
- Equipment and materials
- Work area, level table with ample surface in room that is clean, well-lighted (100 foot-candles at working surface) and well-ventilated, and reasonably free of dust and drafts. The microbial density of air in working area, measured in fallout pour plates taken during plating, should not exceed 15 colonies/plate during 15 min exposure.
- Storage space, free of dust and insects and adequate for protection of equipment and supplies
- Petri dishes, glass or plastic (at least 15 × 90 mm)
- Pipets with pipet aids (no mouth pipetting) or pipettors, 1, 5, and 10 ml, graduated in 0.1 ml units
- Dilution bottles, 6 oz (160 ml), borosilicate-resistant glass, with rubber stoppers or plastic screw caps
- Pipet and petri dish containers, adequate for protection
- Circulating water bath, for tempering agar, thermostatically controlled to 45 ± 1°C
- Incubator, 35 ± 1°C; milk, 32 ± 1°C
- Colony counter, dark-field, Quebec, or equivalent, with suitable light source and grid plate
- Tally register
- Dilution blanks, 90 ± 1 ml Butterfield's phosphate-buffered dilution water (R11); milk, 99 ± 2 ml
- Plate count agar (standard methods) (M124)
- Refrigerator, to cool and maintain samples at 0-5°C; milk, 0-4.4°C
- Freezer, to maintain frozen samples from -15 to -20°C
- Thermometers (mercury) appropriate range; accuracy checked with a thermometer certified by the National Institute of Standards and Technology (NIST)
- Procedure for analysis of frozen, chilled, precooked, or prepared foods
Using separate sterile pipets, prepare decimal dilutions of 10-2, 10-3, 10-4, and others as appropriate, of food homogenate (see Chapter 1 for sample preparation) by transferring 10 ml of previous dilution to 90 ml of diluent. Avoid sampling foam. Shake all dilutions 25 times in 30 cm (1 ft) arc within 7 s. Pipet 1 ml of each dilution into separate, duplicate, appropriately marked petri dishes. Reshake dilution bottle 25 times in 30 cm arc within 7 s if it stands more than 3 min before it is pipetted into petri dish. Add 12-15 ml plate count agar (cooled to 45 ± 1°C) to each plate within 15 min of original dilution. For milk samples, pour an agar control, pour a dilution water control and pipet water for a pipet control. Add agar to the latter two for each series of samples. Add agar immediately to petri dishes when sample diluent contains hygroscopic materials, e.g., flour and starch. Pour agar and dilution water control plates for each series of samples. Immediately mix sample dilutions and agar medium thoroughly and uniformly by alternate rotation and back-and-forth motion of plates on flat level surface. Let agar solidify. Invert solidified petri dishes, and incubate promptly for 48 ± 2 h at 35°C. Do not stack plates when pouring agar or when agar is solidifying.
- Guidelines for calculating and reporting APCs in uncommon cases
Official Methods of Analysis (3) does not provide guidelines for counting and reporting plate counts, whereas Standard Methods for the Examination of Dairy Products, 16th ed. (2) presents detailed guidelines; for uniformity, therefore, use APHA guidelines as modified (6,8). Report all aerobic plate counts (2) computed from duplicate plates. For milk samples, report all aerobic plate (2) counts computed from duplicate plates containing less than 25 colonies as less than 25 estimated count. Report all aerobic plate counts (2) computed from duplicate plates containing more than 250 colonies as estimated counts. Counts outside the normal 25-250 range may give erroneous indications of the actual bacterial composition of the sample. Dilution factors may exaggerate low counts (less than 25), and crowded plates (greater than 250) may be difficult to count or may inhibit the growth of some bacteria, resulting in a low count. Report counts less than 25 or more than 250 colonies as estimated aerobic plate counts (EAPC). Use the following guide:
- Normal plates (25-250). Select spreader-free plate(s). Count all colony forming units (CFU), including those of pinpoint size, on selected plate(s). Record dilution(s) used and total number of colonies counted.
- Plates with more than 250 colonies. When number of CFU per plate exceeds 250, for all dilutions, record the counts as too numerous to count (TNTC) for all but the plate closest to 250, and count CFU in those portions of plate that are representative of colony distribution. See ref. 2 for detailed guidelines. Mark calculated APC with EAPC to denote that it was estimated from counts outside 25-250 per plate range (see D-3).
- Spreaders. Spreading colonies are usually of 3 distinct types: 1) a chain of colonies, not too distinctly separated, that appears to be caused by disintegration of a bacterial clump; 2) one that develops in film of water between agar and bottom of dish; and 3) one that forms in film of water at edge or on surface of agar. If plates prepared from sample have excessive spreader growth so that (a) area covered by spreaders, including total area of repressed growth, exceeds 50% of plate area, or (b) area of repressed growth exceeds 25% of plate area, report plates as spreaders. When it is necessary to count plates containing spreaders not eliminated by (a) or (b) above, count each of the 3 distinct spreader types as one source. For the first type, if only one chain exists, count it as a single colony. If one or more chains appear to originate from separate sources, count each source as one colony. Do not count each individual growth in such chains as a separate colony. Types 2 and 3 usually result in distinct colonies and are counted as such. Combine the spreader count and the colony count to compute the APC.
- Plates with no CFU. When plates from all dilutions have no colonies, report APC as less than 1 times the corresponding lowest dilution used. Mark calculated APC with asterisk to denote that it was estimated from counts outside the 25-250 per plate range. When plate(s) from a sample are known to be contaminated or otherwise unsatisfactory, record the result(s) as laboratory accident (LA).
-
Computing and recording counts (see refs 6, 8)
To avoid creating a fictitious impression of precision and accuracy when computing APC, report only the first two significant digits. Round off to two significant figures only at the time of conversion to SPC. For milk samples, when plates for all dilutions have no colonies, report APC as less than 25 colonies estimated count. Round by raising the second digit to the next highest number when the third digit is 6, 7, 8, or 9 and use zeros for each successive digit toward the right from the second digit. Round down when the third digit is 1, 2, 3, or 4. When the third digit is 5, round up when the second digit is odd and round down when the second digit is even.
Examples
Calculated Count APC 12,700 13,000 12,400 12,000 15,500 16,000 14,500 14,000 - Plates with 25-250 CFU.
- Calculate the APC as follows:
where N = Number of colonies per ml or g of product
Σ c = Sum of all colonies on all plates counted
n1 = Number of plates in first dilution counted
n2 = Number of plates in second dilution counted
d = Dilution from which the first counts were obtainedExample
= 537/0.0221:100 1:1000 232, 244 33, 28
= 24,409
≈ 24,000 - When counts of duplicate plates fall within and without the 25-250 colony range, use only those counts that fall within this range.
- Calculate the APC as follows:
-
All plates with fewer than 25 CFU. When plates from both dilutions yield fewer than 25 CFU each, record actual plate count but record the count as less than 25 × 1/d when d is the dilution factor for the dilution from which the first counts were obtained.
Example
Colonies 1:100 1:1000 EAPC/ml (g) 18 2 <2,500 0 0 <2,500 -
All plates with more than 250 CFU. When plates from both 2 dilutions yield more than 250 CFU each (but fewer than 100/cm2), estimate the aerobic counts from the plates (EAPC) nearest 250 and multiply by the dilution.
Example
Colonies 1:100 1:1000 EAPC/ml (g) TNTC 640 640,000 TNTC, too numerous to count.
EAPC, estimated aerobic plate count. -
All plates with spreaders and/or laboratory accident. Report respectively as Spreader (SPR), or Laboratory Accident (LA).
-
All plates with more than an average of 100 CFU per sq cm. Estimate the APC as greater than 100 times the highest dilution plated, times the area of the plate. The examples below have an average count of 110 per sq cm.
Example
Colonies/Dilution 1:100 1:1000 EAPC/ml (g) TNTC 7,150(a) >6,500,000 EAPC(b) TNTC 6,490 >5,900,000 EAPC a Based on plate area of 65 cm2
b EAPC, estimated APC
c Based on plate area of 59 cm2
- Plates with 25-250 CFU.
Spiral Plate Method
The spiral plate count (SPLC) method for microorganisms in milk, foods, and cosmetics is an official method of the APHA (2) and the AOAC (3). In this method, a mechanical plater inoculates a rotating agar plate with liquid sample. The sample volume dispensed decreases as the dispensing stylus moves from the center to the edge of the rotating plate. The microbial concentration is determined by counting the colonies on a part of the petri dish where they are easily countable and dividing this count by the appropriate volume. One inoculation determines microbial densities between 500 and 500,000 microorganisms/ml. Additional dilutions may be made for suspected high microbial concentrations.
- Equipment and materials
- Spiral plater (Spiral Systems Instruments, Inc., 7830 Old Georgetown Road, Bethesda, MD 20814)
- Spiral colony counter (Spiral Systems) with special grid for relating deposited sample volumes to specific portions of petri dishes
- Vacuum trap for disposal of liquids (2-4 liter vacuum bottle to act as vacuum reservoir and vacuum source of 50-60 cm Hg)
- Disposable micro beakers, 5 ml
- Petri dishes, plastic or glass, 150 × 15 mm or 100 × 15 mm
- Plate count agar (standard methods) (M124)
- Calculator (optional), inexpensive electronic hand calculator is recommended
- Polyethylene bags for storing prepared plates
- Commercial sodium hypochlorite solution, about 5% NaOCl (bleach)
- Sterile dilution water
- Syringe, with Luer tip for obstructions in stylus; capacity not critical
- Work area, storage space, refrigerator, thermometers, tally, incubator, as described for Conventional Plate Count Method, above.
- Sodium hypochlorite solution (5.25%). Available commercially.
- Preparation of agar plates.
Automatic dispenser with sterile delivery system is recommended to prepare agar plates. Agar volume dispensed into plates is reproducible and contamination rate is low compared to hand-pouring of agar in open laboratory. When possible, use laminar air flow hood along with automated dispenser. Pour same quantity of agar into all plates so that same height of agar will be presented to spiral plater stylus tip to maintain contact angle. Agar plates should be level during cooling.
The following method is suggested for prepouring agar plates: Use automatic dispenser or pour constant amount (about 15 ml/100 mm plate; 50 ml/150 mm plate) of sterile agar at 60-70°C into each petri dish. Let agar solidify on level surface with poured plates stacked no higher than 10 dishes. Place solidified agar plates in polyethylene bags, close with ties or heat-sealer, and store inverted at 0-4.4°C. Bring prepoured plates to room temperature before inoculation.
- Preparation of samples.
As described in Chapter 1, select that part of sample with smallest amount of connective tissues or fat globules.
- Description of spiral plater.
Spiral plater inoculates surface of prepared agar plate to permit enumeration of microorganisms in solutions containing between 500 and 500,000 microorganisms per ml. Operator with minimum training can inoculate 50 plates per h. Within range stated, dilution bottles or pipets and other auxiliary equipment are not required. Required bench space is minimal, and time to check instrument alignment is less than 2 min. Plater deposits decreasing amount of sample in Archimedean spiral on surface of prepoured agar plate. Volume of sample on any portion of plate is known. After incubation, colonies appear along line of spiral. If colonies on a portion of plate are sufficiently spaced from each other, count them on special grid which associates a calibrated volume with each area. Estimate number of microorganisms in sample by dividing number of colonies in a defined area by volume contained in same area. Studies have shown the method to be proficient not only with milk (4) but also with other foods (7,10).
- Plating procedure
Check stylus tip angle daily and adjust if necessary. (Use vacuum to hold microscope cover slip against face of stylus tip; if cover slip plane is parallel at about l mm from surface of platform, tip is properly oriented). Liquids are moved through system by vacuum. Clean stylus tip by rinsing for 1 s with sodium hypochlorite solution followed by sterile dilution water for 1 s before sample introduction. This rinse procedure between processing of each sample minimizes cross-contamination. After rinsing, draw sample into tip of Teflon tubing by vacuum applied to 2-way valve. When tubing and syringe are filled with sample, close valve attached to syringe. Place agar plate on platform, place stylus tip on agar surface, and start motor. During inoculation, label petri plate lid. After agar has been inoculated, stylus lifts from agar surface and spiral plater automatically stops. Remove inoculated plate from platform and cover it. Move stylus back to starting position. Vacuum-rinse system with hypochlorite and water, and then introduce new sample. Invert plates and promptly place them in incubator for 48 ± 3 h at 35 ± 1°C.
- Sterility controls
Check sterility of spiral plater for each series of samples by plating sterile dilution water. CAUTION: Prepoured plates should not be contaminated by a surface colony or be below room temperature (water can well-up from agar). They should not be excessively dry, as indicated by large wrinkles or glazed appearance. They should not have water droplets on surface of agar or differences greater than 2 mm in agar depth, and they should not be stored at 0-4.4°C for longer than l month. Reduced flow rate through tubing indicates obstructions or material in system. To clear obstructions, remove valve from syringe, insert hand-held syringe with Luer fitting containing water, and apply pressure. Use alcohol rinse to remove residual material adhering to walls of system. Dissolve accumulated residue with chromic acid. Rinse well after cleaning.
- Counting grid
- Description. Use same counting grid for both 100 and 150 mm petri dishes. A mask is supplied for use with 100 mm dishes. Counting grid is divided into 8 equal wedges; each wedge is divided by 4 arcs labeled l, 2, 3, and 4 from outside grid edge. Other lines within these arcs are added for ease of counting. A segment is the area between 2 arc lines within a wedge. Number of areas counted (e.g., 3) means number of segments counted within a wedge. Spiral plater deposits sample on agar plate in the same way each time. The grid relates colonies on spiral plate to the volume in which they were contained. When colonies are counted with grid, sample volume becomes greater as counting starts at outside edge of plate and proceeds toward center of plate.
- Calibration. The volume of sample represented by various parts of the counting grid is shown in operator's manual that accompanies spiral plater. Grid area constants have been checked by the manufacturer and are accurate. To verify these values, prepare 11 bacterial concentrations in range of 106-103 cells/ml by making 1:1 dilutions of bacterial suspension (use a nonspreader). Plate all Incubate both sets of plates for 48 ± 3 h at 35 ± 1°C. Calculate concentrations for each dilution. Count spiral plates over grid surface, using counting rule of 20 (described in H, below), and record number of colonies counted and grid area over which they were counted. Each spiral colony count for a particular grid area, divided by aerobic count/ml for corresponding spirally plated bacterial concentrations, indicates volume deposited on that particular grid area. Use the following formula:
To check total volume dispensed by spiral plater, weigh amount dispensed from stylus tip. Collect in tared 5 ml plastic beaker and weigh on analytical balance (± 0.2 mg).
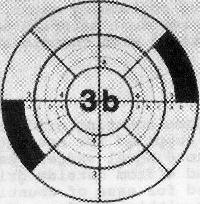
Figure 1. 10 cm plate, area (3b)
- Examination and reporting of spiral plate counts.
Counting rule of 20. After incubation, center spiral plate over grid by adjusting holding arms on viewer. Choose any wedge and begin counting colonies from outer edge of first segment toward center until 20 colonies have been counted. Complete by counting remaining colonies in segment where 20th colony occurs. In this counting procedure, numbers such as 3b, 4c (Fig. l) refer to area segments from outer edge of wedge to designated arc line. Any count irregularities in sample composition are controlled by counting the same segments in the opposite wedge and recording results. Example of spirally inoculated plate (Fig. l) demonstrates method for determining microbial count. Two segments of each wedge were counted on opposite sides of plate with 31 and 30 colonies, respectively. The sample volume contained in the darkened segments is 0.0015 ml. To estimate number of microorganisms, divide count by volume contained in all segments counted. See example under Fig. l.
If 20 CFU are not within the 4 segments of the wedge, count CFU on entire plate. If the number of colonies exceeds 75 in second, third, or fourth segment, which also contains the 20th colony, the estimated number of microorganisms will generally be low because of coincidence error associated with crowding of colonies. In this case, count each circumferentially adjacent segment in all 8 wedges, counting at least 50 colonies, e.g., if the first 2 segments of a wedge contain 19 colonies and the third segment contains the 20th and 76th (or more), count colonies in all circumferentially adjacent first and second segments in all 8 wedges. Calculate contained volume in counted segments of wedges and divide into number of colonies.
When fewer than 20 colonies are counted on the total plate, report results as "less than 500 estimated SPLC per ml." If colony count exceeds 75 in first segment of wedge, report results as "greater than 500,000 estimated SPLC per ml." Do not count spiral plates with irregular distribution of colonies caused by dispensing errors. Report results of such plates as laboratory accident (LA). If spreader covers entire plate, discard plate. If spreader covers half of plate area, count only those colonies that are well distributed in spreader-free areas.
Compute SPLC unless restricted by detection of inhibitory substances in sample, excessive spreader growth, or laboratory accidents. Round off counts as described in I-D, above. Report counts as SPLC or estimated SPLC per ml.
References
- American Public Health Association. 1984. Compendium of Methods for the Microbiological Examination of Foods, 2nd ed. APHA, Washington, DC
- American Public Health Association. 1993. Standard Methods for the Examination of Dairy Products, 16th ed. APHA, Washington, DC.
- Association of Official Analytical Chemists. 1990. Official Methods of Analysis, 15th ed. AOAC, Arlington, VA.
- Donnelly, C.B., J.E. Gilchrist, J.T. Peeler, and J.E. Campbell. 1976. Spiral plate count method for the examination of raw and pasteurized milk. Appl. Environ. Microbiol. 32:21-27.
- Gilchrist, J.E., C.B. Donnelly, J.T. Peeler, and J.E. Campbell. 1977. Collaborative study comparing the spiral plate and aerobic plate count methods. J. Assoc. Off. Anal. Chem. 60:807-812.
- International Dairy Federation. 1987. Milk and Milk Products: Enumeration of Microorganisms—Colony Count at 3°C. Provisional IDF Standard 100A. IDF, Brussels, Belgium.
- Jarvis, B., V.H. Lach, and J.M. Wood. 1977. Evaluation of the spiral plate maker for the enumeration of microorganisms in foods. J. Appl. Bacteriol. 43:149-157.
- Niemela, S. 1983. Statistical evaluation of Results from Quantitative Microbiological Examinations. Report No. 1, 2nd ed. Nordic Committee in Food Analysis, Uppsala, Sweden.
- Tomasiewicz, D.M., D.K. Hotchkiss, G.W. Reinbold, R.B. Read, Jr., and P.A. Hartman. 1980. The most suitable number of colonies on plates for counting. J. Food Prot. 43:282-286.
- Zipkes, M.R., J.E. Gilchrist, and J.T. Peeler. 1981. Comparison of yeast and mold counts by spiral, pour, and streak plate methods. J. Assoc. Off. Anal. Chem. 64:1465-1469.
Hypertext Source: Bacteriological Analytical Manual, Edition 8, Revision A, 1998. Chapter 3.