melanoma
New 3D Atlas of Colorectal Cancer Promises Improved Diagnosis, Treatment
Posted on by Lawrence Tabak, D.D.S., Ph.D.

This year, too many Americans will go to the doctor for tissue biopsies to find out if they have cancer. Highly trained pathologists will examine the biopsies under a microscope for unusual cells that show the telltale physical features of a suspected cancer. As informative as the pathology will be for considering the road ahead, it would be even more helpful if pathologists had the tools to look widely inside cells for the actual molecules giving rise to the tumor.
Working this “molecular information” into the pathology report would bring greater diagnostic precision, drilling down to the actual biology driving the growth of the tumor. It also would help doctors to match the right treatments to a patient’s tumor and not waste time on drugs that will be ineffective.
That’s why researchers have been busy building the needed tools and also mapping out molecular atlases of common cancers. These atlases, really a series of 3D spatial maps detailing various biological features within the tumor, keep getting better all the time. That includes the comprehensive atlas of colorectal cancer just published in the journal Cell [1].
This colorectal atlas comes from an NIH-supported team led by Sandro Santagata, Brigham and Women’s Hospital, Boston, and Peter Sorger, Harvard Medical School, Cambridge, MA, in collaboration with investigators at Vanderbilt University, Nashville, TN. The colorectal atlas joins their previously published high-definition map of melanoma [2], and both are part of the Human Tumor Atlas Network that’s supported by NIH’s National Cancer Institute.
What’s so interesting with the colorectal atlas is the team combined traditional pathology with a sophisticated technique for imaging single cells, enabling them to capture their fine molecular details in an unprecedented way.
They did it using a cutting-edge technique known as cyclic immunofluorescence, or CyCIF. In CyCIF, researchers use many rounds of highly detailed molecular imaging on each tissue sample to generate a rich collection of molecular-level data, cell by cell. Altogether, the researchers captured this fine-scale visual information for nearly 100 million cancer cells isolated from tumor samples representing 93 individuals diagnosed with colorectal cancer.
With this single-cell information in hand, they next created detailed 2D maps covering the length and breadth of large portions of the colorectal cancers under study. Finally, with the aid of first author Jia-Ren Lin, also at Harvard Medical School, and colleagues they stitched together their 2D maps to produce detailed 3D reconstructions showing the length, breadth, and height of the tumors.
This more detailed view of colorectal cancer has allowed the team to explore differences between normal and tumor tissues, as well as variations within an individual tumor. In fact, they’ve uncovered physical features that had never been discovered.
For instance, an individual tumor has regions populated with malignant cells, while other areas look less affected by the cancer. In between are transitional areas that correspond to molecular gradients of information. With this high-resolution map as their guide, researchers can now study what this all might mean for the diagnosis, treatment, and prognosis of colorectal cancer.
The atlas also shows that the presence of immune cells varies dramatically within a single tumor. That’s an important discovery because of its potential implications for immunotherapies, in which treatments aim to unleash the immune system in the fight against cancer.
The maps also provide new insights into tumor structure. For example, scientists had previously identified what they thought were 2D pools of a mucus-like substance called mucin with clusters of cancer cells suspended inside. However, the new 3D reconstruction make clear that these aren’t simple mucin pools. Rather, they are cross sections of larger intricate caverns of mucin interconnected by channels, into which cancer cells make finger-like projections.
The good news is the researchers already are helping to bring these methods into the cancer clinic. They also hope to train other scientists to build their own cancer atlases and grow the collection even more.
In the meantime, the team will refine its 3D tumor reconstructions by integrating new imaging technologies and even more data into their maps. It also will map many more colorectal cancer samples to capture the diversity of their basic biology. Also of note, having created atlases for melanoma and colorectal cancer, the team has plans to tackle breast and brain cancers next.
Let me close by saying, if you’re between the ages of 45 and 75, don’t forget to stay up to date on your colorectal cancer screenings. These tests are very good, and they could save your life.
References:
[1] Multiplexed 3D atlas of state transitions and immune interaction in colorectal cancer. Lin JR, Wang S, Coy S, Chen YA, Yapp C, Tyler M, Nariya MK, Heiser CN, Lau KS, Santagata S, Sorger PK. Cell. 2023 Jan 19;186(2):363-381.e19.
[2] The spatial landscape of progression and immunoediting in primary melanoma at single-cell resolution. Nirmal AJ, Maliga Z, Vallius T, Quattrochi B, Chen AA, Jacobson CA, Pelletier RJ, Yapp C, Arias-Camison R, Chen YA, Lian CG, Murphy GF, Santagata S, Sorger PK. Cancer Discov. 2022 Jun 2;12(6):1518-1541.
Links:
Colorectal Cancer (National Cancer Institute/NIH)
Human Tumor Atlas Network (NCI)
CyCIF-Cyclic Immunofluorescence (Harvard Medical School, Cambridge, MA)
Sandro Santagata (Brigham and Women’s Hospital, Boston)
Peter Sorger (Harvard Medical School)
Jia-Ren Lin (Harvard Medical School)
NIH Support: National Cancer Institute; National Institute of General Medical Sciences; National Institute of Diabetes and Digestive and Kidney Diseases
Fighting Cancer with Next-Gen Cell Engineering
Posted on by Dr. Francis Collins

Researchers continue to make progress with cancer immunotherapy, a type of treatment that harnesses the body’s own immune cells to attack cancer. But Kole Roybal wants to help move the field further ahead by engineering patients’ immune cells to detect an even broader range of cancers and then launch customized attacks against them.
With an eye toward developing the next generation of cell-based immunotherapies, this synthetic biologist at University of California, San Francisco, has already innovatively hacked into how certain cells communicate with each other. Now, he and his research team are using a 2018 NIH Director’s New Innovator Award to build upon that progress.
Roybal’s initial inspiration is CAR-T therapy, one of the most advanced immunotherapies to date. In CAR-T therapy, some of a cancer patient’s key immune cells, called T cells, are removed and engineered in a way that they begin to produce new surface proteins called chimeric antigen receptors (CARs). Those receptors allow the cells to recognize and attack cancer cells more effectively. After expanding the number of these engineered T cells in the lab, doctors infuse them back into patients to enhance their immune systems’s ability to seek-and-destroy their cancer.
As helpful as this approach has been for some people with leukemia, lymphoma, and certain other cancers, it has its limitations. For one, CAR-T therapy relies solely on a T cell’s natural activation program, which can be toxic to patients if the immune cells damage healthy tissues. In other patients, the response simply isn’t strong enough to eradicate a cancer.
Roybal realized that redirecting T cells to attack a broader range of cancers would take more than simply engineering the receptors to bind to cancer cells. It also would require sculpting novel immune cell responses once those receptors were triggered.
Roybal found a solution in a new class of lab-made receptors known as Synthetic Notch, or SynNotch, that he and his colleagues have been developing over the last several years [1, 2]. Notch protein receptors play an essential role in developmental pathways and cell-to-cell communication across a wide range of animal species. What Roybal and his colleagues found especially intriguing is the protein receptors’ mode of action is remarkably direct.
When a protein binds the Notch receptor, a portion of the receptor breaks off and heads for the cell nucleus, where it acts as a switch to turn on other genes. They realized that engineering a cancer patient’s immune cells with synthetic SynNotch receptors could offer extraordinary flexibility in customized sensing and response behaviors. What’s more, the receptors could be tailored to respond to a number of user-specified cues outside of a cell.
In his NIH-supported work, Roybal will devise various versions of SynNotch-engineered cells targeting solid tumors that have proven difficult to treat with current cell therapies. He reports that they are currently developing the tools to engineer cells to sense a broad spectrum of cancers, including melanoma, glioblastoma, and pancreatic cancer.
They’re also engineering cells equipped to respond to a tumor by producing a range of immune factors, including antibodies known to unleash the immune system against cancer. He says he’ll also work on adding engineered SynNotch molecules to other immune cell types, not just T cells.
Given the versatility of the approach, Roybal doesn’t plan to stop there. He’s also interested in regenerative medicine and in engineering therapeutic cells to treat autoimmune conditions. I’m looking forward to see just how far these and other next-gen cell therapies will take us.
References:
[1] Engineering Customized Cell Sensing and Response Behaviors Using Synthetic Notch Receptors. Morsut L, Roybal KT, Xiong X, Gordley RM, Coyle SM, Thomson M, Lim WA. Cell. 2016 Feb 11;164(4):780-91.
[2] Engineering T Cells with Customized Therapeutic Response Programs Using Synthetic Notch Receptors. Roybal KT, Williams JZ, Morsut L, Rupp LJ, Kolinko I, Choe JH, Walker WJ, McNally KA, Lim WA. Cell. 2016 Oct 6;167(2):419-432.e16.
Links:
Car-T Cells: Engineering Patients’ Immune Cells to Treat Cancers (National Cancer Institute/NIH)
Synthetic Biology for Technology Development (National Institute of Biomedical Imaging and Bioengineering/NIH)
Roybal Lab (University of California, San Francisco)
Roybal Project Information (NIH RePORTER)
NIH Support: Common Fund; National Cancer Institute
New Target for Cancer Immunotherapy: Exosomes
Posted on by Dr. Francis Collins

It was once a central tenet of biology that RNA molecules did their work inside the cell. But it’s now clear that RNA molecules are also active outside the cell, with potentially major implications for our health. To learn more about these unrecognized roles, the NIH Common Fund has launched the Extracellular RNA (exRNA) Communication Program.
This month, members of this research consortium described their latest progress in unraveling the secrets of exRNA in a group of 18 papers in the Cell family of journals. And it’s not just RNA that the consortium is studying, it’s also proteins. Among the many exciting results just published is the serendipitous discovery that proteins carried inside tiny, bubble-like vesicles, called exosomes, may influence a cancer’s response to immunotherapy [1]. The work sheds light on why certain cancers are resistant to immunotherapy and points to new strategies for unleashing the immune system in the fight against cancer.
The new findings center on a type of immunotherapy drugs known as checkpoint inhibitors. They are monoclonal antibodies produced by industry that can boost the immune system’s ability to attack and treat cancer.
One of those antibodies specifically targets a protein, called PD-1, on the surface of certain immune cells. When PD-1 binds a similarly named protein, called PD-L1, on the surface of another cell, the interaction prevents immune cells from attacking. Some tumors seem to have learned this and load up on PD-L1 to evade the immune system.
That’s where checkpoint inhibitors come in. By blocking the interaction between PD-1 and PD-L1, the treatment removes a key check on the immune system, allowing certain immune cells to wake up and attack the tumor.
Checkpoint inhibitors work better in some cancer types than in others. In melanoma, for example, up to about 30 percent of patients respond to checkpoint inhibitor therapy. But in prostate cancer, response rates are in the single digits.
Researchers led by Robert Blelloch, a member of the exRNA consortium and a scientist at the University of California, San Francisco, wanted to know why. He and his team looked for clues in RNA within the cells taken from immunotherapy-resistant prostate cancers.
As published in Cell, the researchers got their first hint of something biologically intriguing in an apparent discrepancy in their data. As they expected from prior work, PD-L1 protein was present in the treatment-resistant cancers. But the PD-L1 messenger RNAs (mRNA), which serve as templates for producing the protein, told an unexpected story. The resistant cancer cells made far more PD-L1 mRNAs than needed to produce the modest levels of PD-L1 proteins detected inside the cells.
Where was the missing PD-L1? Blelloch’s team found it in exosomes. The cancer cells were packaging large quantities of the protein inside exosomes and secreting them out of the cell to other parts of the body.
In additional studies with a mouse model of prostate cancer, the researchers found that those PD-L1-packed exosomes travel through the blood and lymphatic systems to lymph nodes, the sites where immune cells become activated. Once there, PD-L1-laden exosomes put the immune system to sleep, preventing certain key cells from locating and attacking the cancer, including the primary tumor and places where it may have spread.
In important follow up studies, the researchers edited two genes in cancer cells to prevent them from producing exosomes. And, in the absence of exosomes, the cells no longer formed tumors. Importantly, both edited and unedited cells still produced PD-L1, but only those that exported PD-L1 in exosomes disarmed the immune system. Studies in a mouse model of immunotherapy-resistant colorectal cancer yielded similar results.
The new evidence suggests that blocking the release of PD-L1 in exosomes, even temporarily, might allow the immune system to launch a successful and sustained attack against a cancer.
Blelloch notes that many intriguing questions remain. For example, it’s not yet clear why antibodies that target PD-L1 on cancer cells don’t disable PD-L1 found in exosomes. The good news is that the new findings suggest it may be possible to find small molecules that do target PD-L1-packed exosomes, unleashing the immune system against cancers that don’t respond to existing checkpoint inhibitors. In fact, Blelloch’s team is already screening for small molecules that might fit the bill.
Since its launch about five years ago, the exRNA Communication Program has published an impressive 480 peer-reviewed papers, including the latest work in the Cell family of journals. I’d encourage readers to click on some of the other excellent work. I hear that another batch of papers will be published later this year.
Reference:
[1] Suppression of exosomal PD-L induces systemic anti-tumor immunity and memory. Poggio M, Hu T, Pai CC, Chu B, Belair CD, Chang A, Montabana E, Lang UE, Fu Q, Fong L, Blelloch R. Cell. 2019 Apr 4;177(2):414-427.
Links:
Video: Unlocking the Mysteries of RNA Communication (Common Fund/NIH)
Immunotherapy to Treat Cancer (National Cancer Institute/NIH)
Blelloch Lab (University of California, San Francisco)
NIH Support: Common Fund; National Cancer Institute; National Center for Advancing Translational Sciences; National Heart, Lung, and Blood Institute; National Institute on Drug Abuse
Watching Cancer Cells Play Ball
Posted on by Dr. Francis Collins
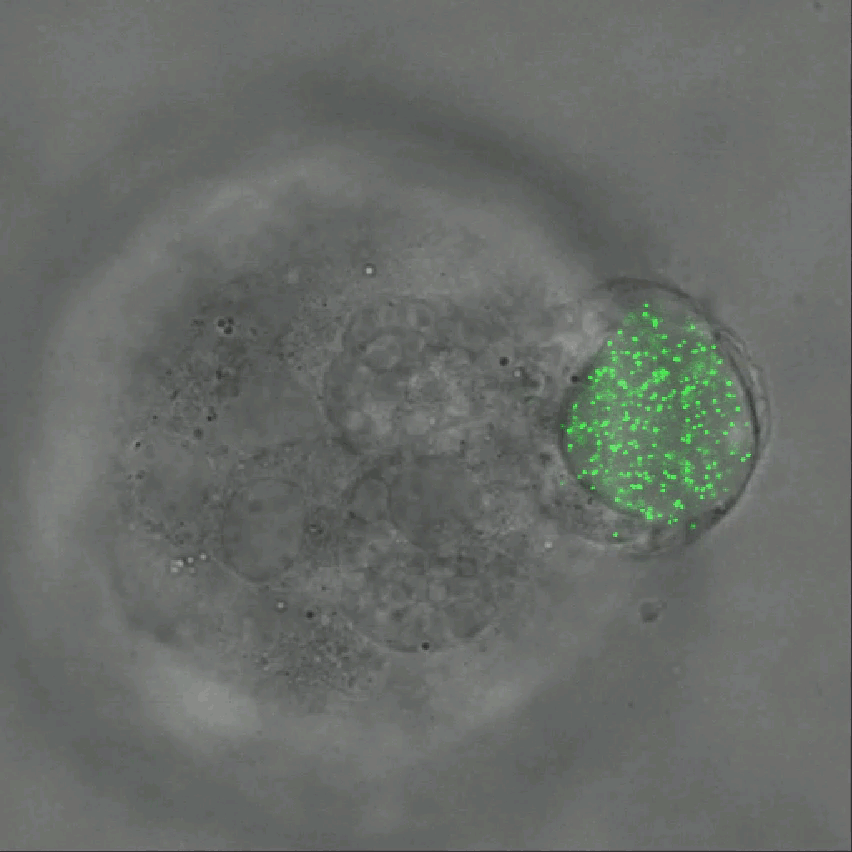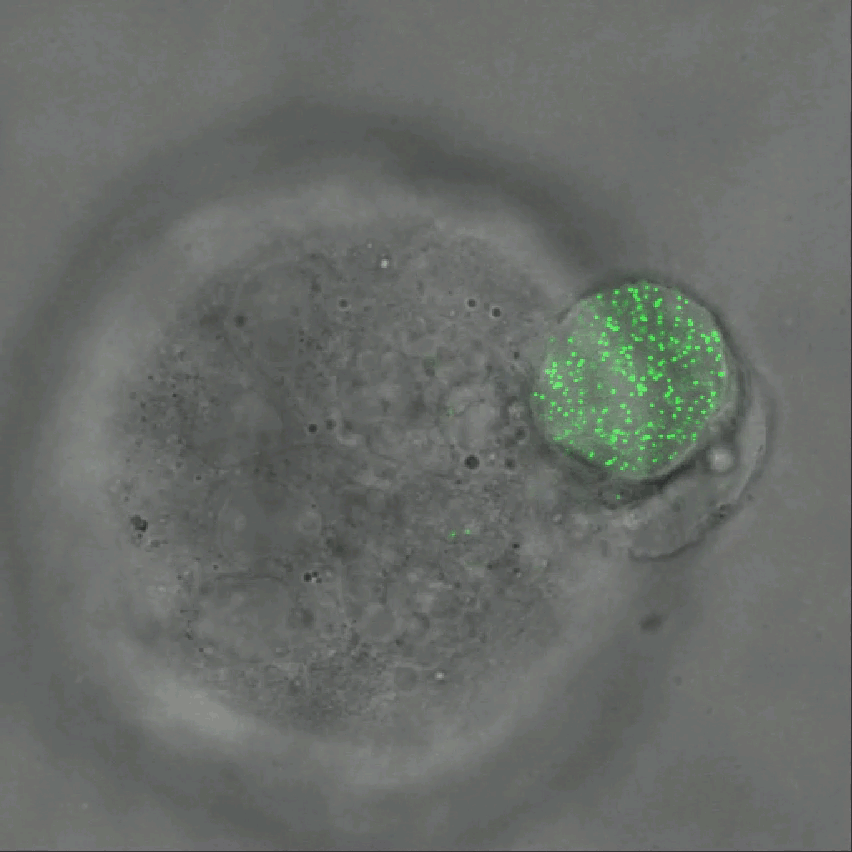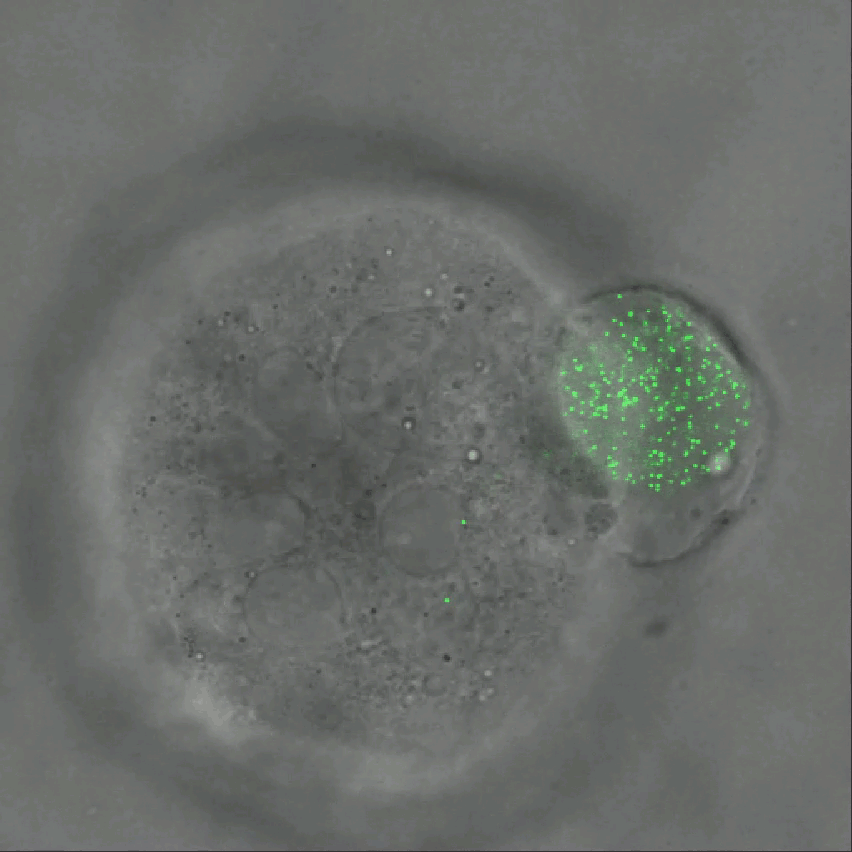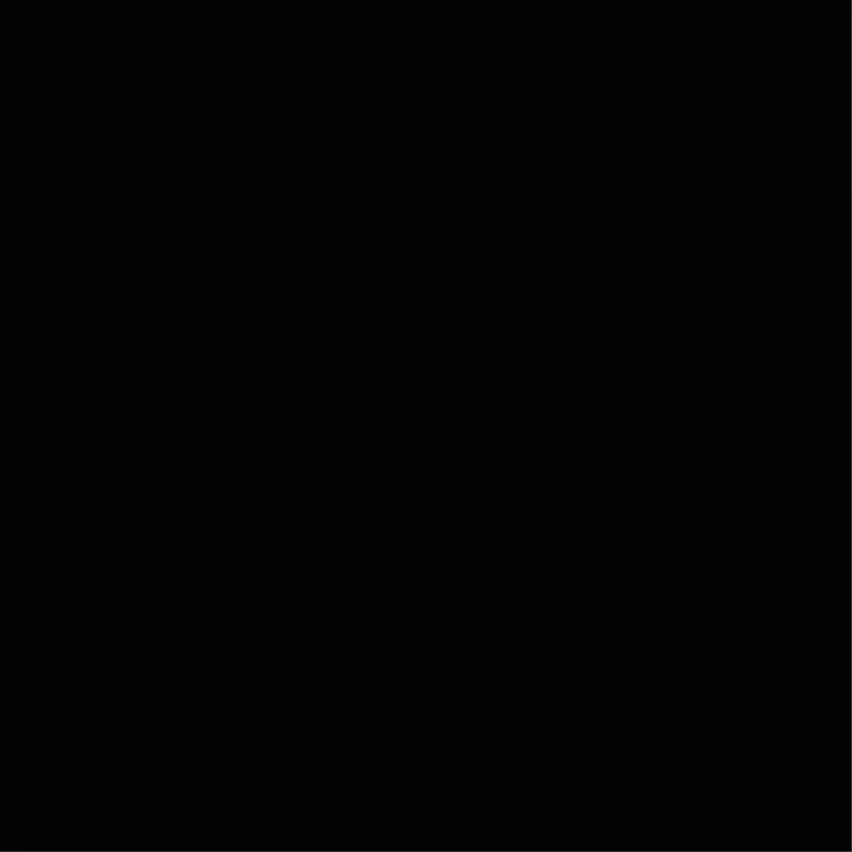
Credit: Ning Wang, University of Illinois at Urbana-Champaign
As tumor cells divide and grow, they push, pull, and squeeze one another. While scientists have suspected those mechanical stresses may play important roles in cancer, it’s been tough to figure out how. That’s in large part because there hadn’t been a good way to measure those forces within a tissue. Now, there is.
As described in Nature Communications, an NIH-funded research team has developed a technique for measuring those subtle mechanical forces in cancer and also during development [1]. Their ingenious approach is called the elastic round microgel (ERMG) method. It relies on round elastic microspheres—similar to miniature basketballs, only filled with fluorescent nanoparticles in place of air. In the time-lapse video above, you see growing and dividing melanoma cancer cells as they squeeze and spin one of those cell-sized “balls” over the course of 24 hours.
Optimizing Radio-Immunotherapy for Cancer
Posted on by Dr. Francis Collins

Zachary Morris
Credit: Alan Leon
Zachary Morris has certainly done some memorable things. As a Rhodes Scholar, he once attended an evening reception at Buckingham Palace, played a game of pick-up football with former President Bill Clinton, and traveled to South Africa to take a Robben Island Prison tour, led by the late Nelson Mandela. But something the young radiation oncologist did during his medical residency could prove even more momentous. He received a special opportunity from the American Board of Radiology to join others in studying how to pair radiation therapy with the emerging cancer treatment strategy of immunotherapy.
Morris’s studies in animals showed that the two treatments have a unique synergy, generating a sustained tumor-specific immune response that’s more potent than either therapy alone. But getting this combination therapy just right to optimize its cancer-fighting abilities remains complicated. Morris, now a researcher and clinician at the University of Wisconsin School of Medicine and Public Health, Madison, has received a 2017 NIH Director’s Early Independence Award to look deeper into this promising approach. He and his collaborators will use what they learn to better inform their future early stage clinical trials of radio-immunotherapy starting with melanoma, head and neck cancers, and neuroblastoma.
Creative Minds: Applying CRISPR Technology to Cancer Drug Resistance
Posted on by Dr. Francis Collins

Patrick Hsu
As a child, Patrick Hsu once settled a disagreement with his mother over antibacterial wipes by testing them in controlled experiments in the kitchen. When the family moved to Palo Alto, CA, instead of trying out for the football team or asking to borrow the family car like other high school kids might have done, Hsu went knocking on doors of scientists at Stanford University. He found his way into a neuroscience lab, where he gained experience with the fundamental tools of biology and a fascination for understanding how the brain works. But Hsu would soon become impatient with the tools that were available to ask some of the big questions he wanted to study.
As a Salk Helmsley Fellow and principal investigator at the Salk Institute for Biological Studies, La Jolla, CA, Hsu now works at the intersection of bioengineering, genomics, and neuroscience with a DNA editing tool called CRISPR/Cas9 that is revolutionizing the way scientists can ask and answer those big questions. (This blog has previously featured several examples of how this technology is revolutionizing biomedical research.) Hsu has received a 2015 NIH Director’s Early Independence award to adapt CRISPR/Cas9 technology so its use can be extended to that other critically important information-containing nucleic acid—RNA.Specifically, Hsu aims to develop ways to use this new tool to examine the role of a certain type of RNA in cancer drug resistance.
Precision Oncology: Creating a Genomic Guide for Melanoma Therapy
Posted on by Dr. Francis Collins

Caption: Human malignant melanoma cell viewed through a fluorescent, laser-scanning confocal microscope. Invasive structures involved in metastasis appear as greenish-yellow dots, while actin (green) and vinculin (red) are components of the cell’s cytoskeleton.
Credit: Vira V. Artym, National Institute of Dental and Craniofacial Research, NIH
It’s still the case in most medical care systems that cancers are classified mainly by the type of tissue or part of the body in which they arose—lung, brain, breast, colon, pancreas, and so on. But a radical change is underway. Thanks to advances in scientific knowledge and DNA sequencing technology, researchers are identifying the molecular fingerprints of various cancers and using them to divide cancer’s once-broad categories into far more precise types and subtypes. They are also discovering that cancers that arise in totally different parts of the body can sometimes have a lot in common. Not only can molecular analysis refine diagnosis and provide new insights into what’s driving the growth of a specific tumor, it may also point to the treatment strategy with the greatest chance of helping a particular patient.
The latest cancer to undergo such rigorous, comprehensive molecular analysis is malignant melanoma. While melanoma can rarely arise in the eye and a few other parts of the body, this report focused on the more familiar “cutaneous melanoma,” a deadly and increasingly common form of skin cancer [1]. Reporting in the journal Cell [2], The Cancer Genome Atlas (TCGA) Network says it has identified four distinct molecular subtypes of melanoma. In addition, the NIH-funded network identified an immune signature that spans all four subtypes. Together, these achievements establish a much-needed framework that may guide decisions about which targeted drug, immunotherapy, or combination of therapies to try in an individual with melanoma.
Knocking Out Melanoma: Does This Triple Combo Have What It Takes?
Posted on by Dr. Francis Collins
It would be great if we could knock out cancer with a single punch. But the more we learn about cancer’s molecular complexities and the immune system’s response to tumors, the more it appears that we may need a precise combination of blows to defeat a patient’s cancer permanently, with no need for a later rematch. One cancer that provides us with a ringside seat on the powerful potential—and tough challenges—of targeted combination therapy is melanoma, especially the approximately 50% of advanced tumors with a specific “driver” mutation in the BRAF gene [1].
Drugs that target cells carrying BRAF mutations initially provided great hope for melanoma, with many reports of dramatic shrinkage of tumors in patients with advanced disease. But almost invariably, the disease recurred and was no longer responsive to those same drugs. A few years ago, researchers thought they’d come up with a solid combination to fight BRAF-mutant melanoma: a one-two punch that paired a BRAF-inhibiting drug with an agent that sensitized the immune system [2]. However, when that combo was tested in humans, the clinical trial had to be stopped early because of serious liver toxicity [3]. Now, in a mouse study published in Science Translational Medicine, NIH-funded researchers at the University of California, Los Angeles (UCLA) provide renewed hope for a safe, effective combination therapy for melanoma—with a strategy that adds a third drug to the mix [4].
Lessons from a High School Student: Motivation + Perseverance = Success
Posted on by Dr. Francis Collins
It may surprise you to learn that the poised young woman featured in this video was a sophomore in high school at the time the film was made. Today, Emily Ashkin is a high school senior with impressive laboratory experience and science awards to her name. As it happens, she’s also introducing me when I deliver a keynote address at the Melanoma Research Alliance’s annual scientific meeting — today, here in Washington, D.C.
What struck me most when I heard Emily’s story was her fearlessness. When mentoring young students, helping some to believe in themselves can be a real challenge. Not Emily. She faces her challenges by seeking solutions, asking—as she does in the video—“Why can’t that be me?”
Next Page


